 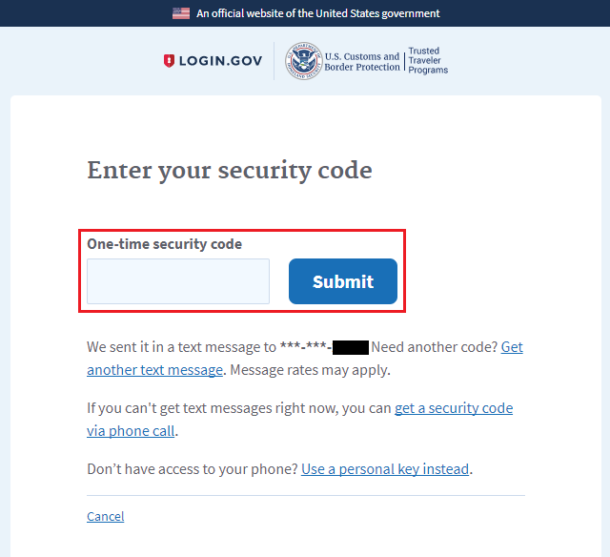
Nuclear speckles were first visualized by transmission EM (TEM) as dense clusters of 20–25-nm-diameter RNP granules ( Fakan and Puvion, 1980) termed interchromatin granule clusters, and they have alternatively been proposed to be storage sites for RNA-processing components ( Spector and Lamond, 2011) or transcription hubs for a subset of active genes ( Xing et al., 1995 Shopland et al., 2003 Hall et al., 2006). Nuclear speckles, excluded from the nuclear periphery and enriched toward the nuclear center ( Carter et al., 1991), are an excellent candidate for such a nuclear compartment. Alternatively, this stochastic radial positioning of genes could be the consequence of a more deterministic positioning of genes relative to a nuclear compartment or compartments that themselves show a stochastic radial positioning. However, the functional significance of this radial positioning has been difficult to rationalize given the large variability of gene positioning within individual nuclei ( Takizawa et al., 2008 Kölbl et al., 2012). Using DNA FISH, a population-based, statistical shift toward the nuclear center has been observed for a number of genes undergoing transcriptional activation ( Takizawa et al., 2008), leading to the proposal of a gradient of increased transcriptional activity from the nuclear periphery to center ( Takizawa et al., 2008 Bickmore, 2013). 
For example, whether transcriptionally active chromosome regions are targeted reproducibly to particular nuclear compartments has been a long-standing question. 
What is needed is an ability to translate microscopic views of DNA position relative to nuclear compartments (such as the nuclear lamina, nucleolus, or nuclear speckles) into genome-wide maps that show how close loci are to a given compartment and how the chromosomal fiber traverses between compartments. However, these 3C (chromosome conformation capture)-based methods do not directly report on chromosome positioning within nuclei. New genomic methods such as Hi-C ( Lieberman-Aiden et al., 2009 Rao et al., 2014) have generated increasing interest in how 3D chromosome folding may regulate genome functions during development or in health and disease. While the human genome has been sequenced, how this linear genome sequence folds in 3D within the nucleus remains largely unknown. Our results demonstrate the capability of TSA-Seq for genome-wide mapping of nuclear structure and suggest a new model for spatial organization of transcription and gene expression. This gradient represents a convolution of multiple spatially separated nuclear domains including two types of transcription “hot zones.” Transcription hot zones protruding furthest into the nuclear interior and positioning deterministically very close to nuclear speckles have higher numbers of total genes, the most highly expressed genes, housekeeping genes, genes with low transcriptional pausing, and super-enhancers. Ensemble-averaged results in K562 cells reveal a clear nuclear lamina to speckle axis correlated with a striking spatial gradient in genome activity. In this study, we describe TSA-Seq, a new mapping method capable of providing a “cytological ruler” for estimating mean chromosomal distances from nuclear speckles genome-wide and for predicting several Mbp chromosome trajectories between nuclear compartments without sophisticated computational modeling. While nuclear compartmentalization is an essential feature of three-dimensional genome organization, no genomic method exists for measuring chromosome distances to defined nuclear structures. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
